Computer can infer rules of the forest
By Anne Ju

A forest full of rabbits and foxes, a bubbling vat of chemical reactants, and complex biochemical circuitry within a cell are, to a computer, similar systems: Many scenarios can play out depending on a fixed set of rules and individual interactions that can’t be precisely predicted – chemicals combining, genes triggering cascades of chemical pathways, or rabbits multiplying or getting eaten.
Predicting possible outcomes from a set of rules that contain uncertain factors is often done using what’s called stochastic prediction. What has eluded scientists for decades is doing the reverse: To find out what the rules were, simply by observing the outcomes.
Researchers led by Hod Lipson, associate professor of mechanical and aerospace engineering and of computing and information science, have published new insight into automated stochastic inference that could help unravel the hidden laws in fields as diverse as molecular biology to population ecology to basic chemistry.
Their study, published online July 22 in Proceedings of the National Academy of Sciences, describes a new computer algorithm that allows machines to infer stochastic reaction models without human intervention, and without any previous knowledge on the nature of the system being modeled.
With their algorithm, Lipson and colleagues have devised a way to take intermittent samples – for example, the number of prey and predating species in a forest once a year, or the concentration of different species in a chemical bath once an hour – and infer the likely reactions that led to that result. They’re working backward from traditional stochastic modeling, which typically uses known reactions to simulate possible outcomes. Here, they’re taking outcomes and coming up with reactions, which is much trickier, they say.
“This could be very useful if you wanted to learn the driving rules for not just foxes and rabbits, but any evolving system with interacting agents,” Lipson said. “There is a whole lot of science that is based on this kind of modeling.”
The researchers, including first author Ishanu Chattopadhyay, a Cornell postdoctoral associate, teamed with Anna Kuchina and Gurol Suel, molecular biologists at University of California, San Diego, to test their algorithms using real data. In one experiment, they applied the algorithm to a set of gene expression measurements of a model bacterium B. subtilis.
They gleaned similar insights by studying the fluctuating numbers of micro-organisms in a closed ecosystem; the algorithm came up with reactions that correctly identified the predators, the prey and the dynamical rules that defined their interactions.
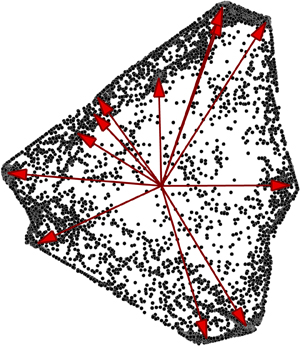
Their key insight was to look at relative changes of the concentration of the interacting agents, irrespective of the time at which such changes were observed. This collective set of relative population updates has some important mathematical properties, which could be related back to the hidden reactions driving the system.
“We figured out that there’s what’s called an invariant geometry, a geometrical feature of the data set that you can uncover even from sparse intermittent samples, without knowing any of the underlying rules,” Chattopadhyay said. “The geometry is a function of the rules, and once you find that out, there is a way to find out what the reactions are.”
The bigger picture in this study is to give scientists better tools for taking massive amounts of data and coming up with simple, insightful explanations, Lipson said.
“This is a tool in a suite of emerging ‘automated science’ tools researchers can use if they have data from some experiment, and they want the computer to help them understand what’s going on – but in the end, it’s the scientist who has to give meaning to these models,” Lipson said.
The study, “Inverse Gillespie: Inferring Stochastic Reaction Mechanisms From Intermittent samples,” was supported by the U.S. Army Research Office Biomathematics program, the National Science Foundation and the Defense Threat Reduction Agency.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe